|
|
Journal of Advanced Veterinary Research Volume 9, Issue 3, 2019, Pages: 102-106 www.advetresearch.com |
|
|
Bacteriological and Molecular Identification of Thermophilic Campylobacters of Animal and Human Origins in Beni-Suef Governorate, Egypt |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Sherin R. Rouby1, Gihan K. Abdel-Latef2, Sahar Abdel Aleem Abdel Aziz2* |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
1Department of Veterinary Medicine, Faculty of Veterinary Medicine, Beni-Suef University, Beni-Suef 62511, Egypt. 2Department of Hygiene, Zoonoses and Epidemiology, Faculty of Veterinary Medicine, Beni-Suef University, Beni-Suef 62511, Egypt. |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Received: 13 June 2019; Accepted: 2 July 2019 *Corresponding author: Sahar Abdel Aleem Abdel Aziz (abdelaziz.sahar@yahoo.com) |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Abstract |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Thermophilic species of the genus Campylobacter are generally considered commensals of livestock and the leading cause of bacterial food-borne zoonoses. The present study was delineated to clarify the role of Campylobacter species as a diarrheagenic pathogen in animals and man and to investigate the fecal carriage rate of Campylobacters in animals and in-contact humans. A total number of 78 fecal samples were collected from diarrheic and non-diarrheic cattle (n=26), sheep (n=28) and humans (n=24). Samples were enriched in Preston broth, followed by streaking on selective Campylobacter agar base medium. The suspected colonies were tested morphologically and biochemically. Campylobacter spp. was recovered from 29 (37.17%) out of 78 fecal samples (34.61%, 42.85% and 33.33%) for cattle, sheep and humans, respectively. Positive correlation between the occurrence of diarrhea and the isolation of Campylobacters was observed in samples of human origin while in adult ruminants particularly sheep, high fecal carriage rate was observed in non-diarrheic animals. The isolates were identified to genus and species levels by polymerase chain reaction targeting the 16S rRNA gene, the mapA gene and the ceuE gene which revealed that all of isolates were Campylobacter jejuni. These findings pose a significant epidemiological implication where cattle and sheep act as vehicles of, and excrete Campylobacter jejuni which is capable of causing disease in the local community in the area of investigation. |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Keywords: Campylobacter, Commensals, Food-borne, Thermophilic, Zoonotic |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Introduction |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
From the beginning of the last century, Campylobacter was recognized as an animal pathogen. Currently the genus Campylobacter counts 27 species, 9 subspecies and 3 biovars, out of them 19 species are considered to be pathogenic for humans and 9 for the animals (Ngulukun, 2017) with the thermotolerant Campylobacter jejuni (C. jejuni) and Campylobacter coli (C. coli) represent the most important species in terms of the food safety point and public health potentials (Foster et al., 2004). Thermotolerant Campylobacters are frequent colonizers of the intestine of livestock such as cattle and sheep (Leuchtefeld and Wang, 1982), which represent potential reservoirs for humans. High fecal carriage rates of C. jejuni/coli have been reported in young animals than in older animals. In the latter, the organisms can be occasionally detected in feces due to low numbers or due to intermittent shedding. Contact with infected animals, consumption of contaminated water or raw milk and travelling in high prevalence areas constitute the major risk factors in human infections (Friedman et al., 2000). Campylobacter jejuni/coli are the leading cause of food-borne bacterial gastroenteritis worldwide (Moore et al., 2005; Behringer et al., 2011; Mdegela et al., 2011; Silva et al., 2011). Campylobacteriosis is associated with 400–500 million cases of diarrhea per year; this number of cases often exceeds those recorded for salmonellosis and shigellosis (Ruiz-Palacios, 2007). In developing countries, the disease remains underreported due to the lack of regular surveillance programs (Coker et al., 2002; Workman et al., 2006). Human infection is characterized by self-limiting diarrhea, acute dysentery, abdominal cramps lasting for 7 days and fever (Skirrow and Blaser, 2000; Moore et al., 2005; Messens et al., 2009, Blanco and pall, 2018) however, some individuals develop sequelae after the acute phase, C. jejuni infection may lead to autoimmune conditions known as Guillain-Barré syndrome (GBS) and Miller Fisher syndrome. They have also been reported to be involved in extra-gastrointestinal manifestations, including bacteremia, lung infections, brain abscesses, meningitis, and reactive arthritis (Man, 2011). More recent studies suggested that C. jejuni infections can lead to inflammatory bowel disease such as Crohn’s disease as well as abortion (Horrocks et al., 2009). Diagnosis of Campylobacteriosis relies on isolation and identification of the causative organism using a battery of biochemical tests. Campylobacters require microaerobic conditions and grow optimally at 42°C; they neither ferment nor oxidize carbohydrates. The close relationship between the species makes the tests unreliable in distinction between Campylobacter species (Presson and Olsen, 2005). Using molecular techniques, Campylobacter can be easily identified not only to genus level but also to species level (Moore et al., 2005; Silva et al., 2011). The objective of the present study was directed to clarify the role of Campylobacter species as a diarrheagenic pathogen in animals and man employing different bacteriological, biochemical and molecular tools. |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Materials and methods |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Area of study The present study was performed on cattle, sheep and in-contact humans located in Beni-Suef Governorate (coordinates: 29°04′N31°05′E), Egypt from June 2016 till May 2017. Animals and clinical samples A total number of 54 animals consisting of 26 cattle (six calves of 1-8 weeks and 20 adults) and 28 sheep (five newly born lambs, 12 sheep of 6-12 months and 11 >one years old) were examined for signs of enteritis in terms of fecal consistency, color and odor. Amongst 54 animals, five cattle (four calves and one adult) and four sheep (one newly born lamb and three sheep of 6-12 months) were suffered clinically from enteritis manifested by diarrhea. Fecal samples of these animals were subjected for exhaustive bacteriological examination for isolation of Campylobacter spp. Human samples Diarrheic (n=7) and non-diarrheic (n=17) humans in actual contact with the above mentioned examined animals were included in this study. Stool samples of humans were subjected for exhaustive bacteriological examination for isolation of Campylobacter spp. Samples were kept at 4°C and processed for isolation of Campylobacter spp. within one hour of collection. Data of animal and human samples are illustrated in Table 1. The present study was approved by the institutional animal care and use committee of Beni-Suef University (BSU-IACUC) and IRB (Faculty of Medicine, Beni-Suef University). Table 1. Data of animal and human samples and results of isolation
Isolation and phenotypic identification of Campylobacter species For selective isolation of Campylobacter spp., approximately one gram from each fecal sample was inoculated into a tube containing 9 ml of Preston Campylobacter selective enrichment broth (prepared by adding 12.5 g of Nutrient Broth No. 2 in 475 ml of distilled water and sterilized by autoclaving followed by cooling at 50°C then adding aseptically 25 ml of lysed horse blood, and one vial of Preston Campylobacter Selective Supplement (Oxoid Ltd, Basingstoke, Hampshire, England). Broth was incubated at 42 °C for 24 hrs. in anaerobic jar using commercial Gas Packs System BBL (5.0% O2, 10.0% CO2 and 85.0% N2, Oxoid, Unipath, Basingstoke, Hampshire, England) (Bolton and Robertson, 1982). A loopful from each broth was streaked on the surface of Campylobacter selective agar base containing an antibiotic supplement for the selective isolation of Campylobacter species (HiMedia Laboratories, Mumbai, India) and 5% (v/v) defibrinated sheep blood. All inoculated plates were incubated for 24-48 hrs. at 42 °C under the above mentioned conditions (Roberts and Greenwood, 2003). Suspected colonies that appeared as translucent white, moist and glistening were picked and re-streaked onto selective media and incubated at 41.5 °C into gas pack apparatus under microaerophilic condition for 24 hrs. for purification. Phenotypic identification of presumptive Campylobacter species was done using standard biochemical and microbiological techniques according to On, (1996). DNA extraction Pure colonies on selective media plates were picked and suspended in sterile deionized distilled water then boiled in a water bath for 10 minutes. The samples were cooled immediately for 5-10 minutes on ice and centrifuged at 10000 × g for 10 min. The supernatants were used as DNA templates for the molecular identification (Van Eys et al., 1989). DNA amplification The 16S rRNA gene (Linton et al., 1997) was amplified to detect Campylobacter on the genus level while; the mapA gene (Stucki et al., 1995) and the ceuE gene (Gonzalez et al., 1997) were selected to detect Campylobacters on species level (C. jejuni and C. coli, respectively). Primer sequences and origin are illustrated in Table 2. Amplification reactions were carried out using a DNA thermal cycler (Labnet® Multigen Gradient thermal cycler, Catalog TC9600-G- 230V (Labnet international, Inc. Edison, NJ, USA) with the following program: one cycle of 5 min at 95 °C, 35 cycles each consisting of 45 s at 94° C, 45 s at 59 °C, 1 min. at 72 °C and a final extension step of 10 min. at 72 °C. Table 2. Primer sequences specific for Campylobacter organisms
DNA electrophoresis The PCR amplicons were analyzed by running 15 µl of the PCR products in 1.5% agarose gel stained with ethidium bromide (0.5µg/ml) and visualized under ultra-violet (UV) light using gel documentation and analysis system. |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Results |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
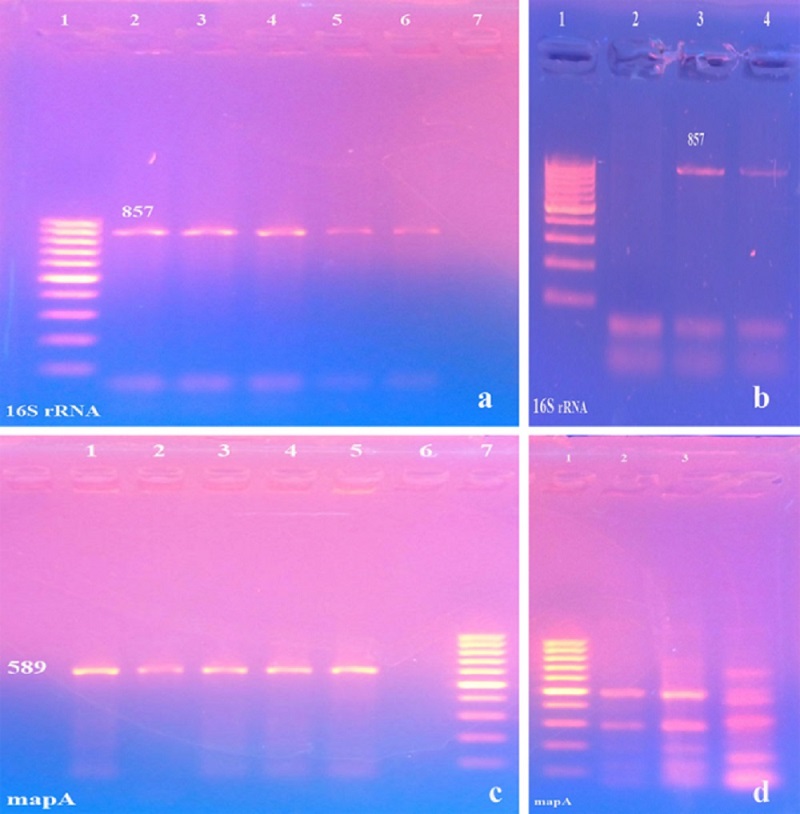
Out of 78 fecal samples screened for the existence of Campylobacters, 29 samples generated typical bacterial colonies (Table 1). Colonies appeared as translucent white, moist and glistening. The highest isolation rates of Campylobacter species were found in sheep (42.85%) followed by cattle (34.61%) and then humans (33.33%). Campylobacters were isolated from diarrheic and non-diarrheic fecal samples; however, in humans and young animal samples there was a strong association between the isolation of Campylobacters and the occurrence of diarrhea (71.43% and 100%, respectively) (Table 1). The 16S rRNA gene based PCR on cultures collected from selective agar plates generated PCR products with a length of 857 bp (Fig. 1a, b) and the mapA gene based PCR generated PCR products with a length of 589 bp in all isolates (Fig. 1c, d), while no results could be obtained using the ceuE gene based PCR, thereby, all isolates were identified as C. jejuni. |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Fig. 1. Gel electrophoresis of PCR product using 16S rRNA gene specific primer (a, b) and mapA gene specific primer for Campylobacter organisms (c, d). a: Lane 1: Ladder 100 bp, Lanes 2,3,4: Sheep isolates, Lanes 5,6: Cattle isolates and Lane 7: control negative. b: Lane 1: Ladder 100 bp, Lane 2: control negative and Lanes 3,4: Human isolates. c: Lanes 1,2,3: Sheep isolates, Lanes 4,5: Cattle isolates and Lane 6: control negative Lane 7: Ladder 100 bp. d: Lane 1: Ladder 100 bp and Lanes 2,3,4: Human isolates. |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Discussion |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Thermotolerant Campylobacter infections constitute a zoonotic and public health problem. In the present study, a total of 54 domestic animals consisted of cattle (n=26) and sheep (n=28) were surveyed for fecal carriage of Campylobacter spp. beside 24 human stool samples from livestock contacts throughout the period between June 2016 and May 2017 in Beni-Suef, Egypt. Out of 78 fecal samples, 29 samples (37.17%) yielded typical bacterial colonies of Campylobacter spp. In the current study, the enrichment step was employed prior to the inoculation and culture technique in order to enhance the isolation rate. Several authors confirmed that the average number of Campylobacter in intestinal samples collected from adult ruminants is low (Stanley et al., 1998; Nielsen, 2002), thereby an enrichment step is necessary to increase the recovery of Campylobacter from bovine and ovine samples. The highest isolation rates were observed in sheep (42.85%), followed by cattle (34.61%). Thermophilic Campylobacters are readily colonize the intestinal tract of ruminants (Stanley and Jones, 2003) however isolation rates vary between herds and flocks as a result of several factors such as husbandry practices, stocking density, age of animal and the housing method (Stanley and Jones, 2003). In the current study high isolation rate was observed in young animals suffered from diarrhea (100%) while in adult animals particularly sheep high isolation rate (63.63) was observed among apparently health animals. Newborn animals acquire Campylobacters horizontally from the farm environment components; these bacteria easily colonize the host due to the underdevelopment of their gastrointestinal tract (Stanley et al., 1998). Association of Campylobacter infection with diarrhea can be explained on the basis of invasion of C. jejuni to the intestine that causes cellular inflammation resulting from the production of cytotoxins followed by the reduction of the absorptive capacity of the intestine as reported by Van Deun et al. (2007) who believed that the ability of this pathogen to reach the intestinal tract is in part due to its resistance to gastric acids and also to bile salts. The obtained results indicate that Campylobacters is frequently isolated from the intestine of healthy and diarrheic animals and considered to be part of the normal intestinal flora, hence domestic ruminants play an integral role in the ecology of C. jejuni and may serve as source of infection resulting in outbreaks of disease or sporadic cases in humans as explained by Di Giannatale et al. (2014). Under such situation of the frequent isolation of C. jejuni as substantiated in this study, it is of at most importance to take in consideration that although C. jejuni mainly colonizes the gastrointestinal tract in animals, it may cross the intestinal epithelial barrier leading to bacteremia that occasionally cause mastitis in cattle and may reach the gravid uterus, resulting in subsequent placentitis, fetal infection and abortion as reported by Di Giannatale et al. (2014). Regarding humans, the highest isolation rate of Campylobacter shedding was obtained from diarrheic patients (71.43%). The obtained results indicate that C. jejuni is one of the most important bacterial causes of diarrhea among people leading to inflammatory enteritis and acute dysentery with severe abdominal pain and fever as the commonest manifestations as reported by Blanco and Pall (2018). The most commonly reported symptoms of patients with laboratory-confirmed infections include diarrhea, fever, and abdominal cramping (Blaser et al., 1983) Less frequently, C. jejuni infections produce bacteremia, septic arthritis, and other extra-intestinal symptoms (Peterson, 1994). Campylobacter is the most common bacterium inducing gastroenteritis in human beings globally in developed and developing countries especially in Africa, Asia and Middle East (Blaser, 1997; Allos, 2001; Dasti et al., 2010; Kaakoush et al., 2015) that may lead to fatal consequence especially in very young children, geriatric people and immune compromised patients (WHO, 2017). According to the bacteriological findings obtained in this study, Campylobacter jejuni is shed in the feces of both diarrheic and healthy animals and humans. However fecal shedding by healthy animals is intermittent and only few organisms are shed (Di Giannatale et al., 2014). Campylobacter isolates isolated in this study could be identified by PCR on both genus and species levels. Employing PCR, all the 29 Campylobacter isolates yielded 857 bp amplicon specific for 16S rRNA gene, and showed the specific 589 bp in mapA gene typical for C. jejuni. C. jejuni had been reported in Egypt as 8.4% (Shimaa et al., 2015 in Cairo, Fayoum Minya and El-Qalubia governorates), 5.2% (Girgis et al., 2014 in Ain Shams University hospital) and 27.5% (Moustafa et al., 2018 in Assiut Unit hospital). Although sodium hippurate hydrolysis reaction is the only biochemical test used to differentiate between C. jejuni and C. coli, the test is time consuming and sometimes difficult to interpret when the enzymatic activity is impaired under the methodological condition (Rautelin et al., 1999). PCR is a recommended molecular technique for a decisive diagnosis and distinction between Campylobacter species (Oyofo et al., 1992; Comi et al., 1995; Sails et al., 1998) and offers an effective alternative to traditional biochemical methods for field studies. The current study elucidates the important role that cattle and sheep play in the dissemination of Campylobacters. Apparently healthy animals shedding Campylobacters in their feces may contribute in the spread of infection amongst the herd and pose a high zoonotic risk to in-contact humans through contamination of milk at the farm level, the carcasses at slaughter house and surface water during removal of abattoir effluents to land (Gannon, 1999). |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Conclusion |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Domestic ruminants play an integral role in the ecology of C. jejuni and may serve as source of infection resulting in outbreaks of disease or sporadic cases in humans. PCR is a highly recommended molecular technique for a decisive diagnosis and distinction between different Campylobacter species recovered from animals and humans. |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Acknowledgments |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
We would like to thank Prof. Hosein I. H. for kindly editing the manuscript. |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Conflict of Interests |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
None of the authors of this paper has a financial or personal relationship with other people or organizations that could inappropriately influence or bias the content of the paper. |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Funding information The present study was not received any specific grants from funding agencies in the public, commercial or not-for profit sectors. |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
References |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Allos, B.M. 2001. C. jejuni infections: Update on emerging issues and trends. Clinical Infectious Disease 32, 1201–1206. Behringer, M.W.G., Miller, O., Oyarzabal, A.M., 2011. Typing of C. jejuni and C. coli isolated from live broilers and retail broiler meat by flaA-RFLP, MLST, PFGE and REP-PCR. Journal Microbiology Methods 84, 194-201. Blanco, D.A., Pall, H., 2018. Principles and Practice of Pediatric Infectious Diseases. (Fifth Edition) Elsevier Inc. pp. 1588-1662. Blaser, M.J., 1997. Epidemiologic and clinical features of C. jejuni infections. Journal Infectious Disease 176(Suppl 2), 103–105. Blaser, M.J., Wells, J.G., Feldman, R.A., Pollard, R.A., Allen, J.R., 1983. The Collaborative Diarrheal Disease Study Group. Campylobacter enteritis in the United States: a multicenter study. Ann. Intern. Med. 98, 360–365. Bolton, F.J., Roberston, L., 1982. A selective medium for isolating C. jejuni/coli. Journal Clinical Pathology 35, 462-467. Comi, G., Ferroni, P., Cocolin, L., Cantoni, C., Manzano, M., 1995. Detection and identification of C. coli and C. jejuni by two-step polymerase chain reaction. Molecular Biotechnology 3, 266-26. Coker, A.O., Isokpehi, R.D., Thomas, B.N., Amisu, K.O., Obi, C.L., 2002. Human Campylobacteriosis in developing countries. Emerging Infectious Diseases 8, 237-243. Dasti, J.I., Tareen, A.M., Lugert, R., Zautner, A.E., Gross, U., 2110. C. jejuni: a brief overview on pathogenicity-associated factors and disease-mediating mechanisms. International Journal Medical Microbiology 300, 205–11. Di Giannatale, E., Di Serafino, G., Zilli, K., Alessiani, A., Sacchini, L., Garofolo, G., 2014. Characterization of antimicrobial resistance patterns and detection of virulence genes in Campylobacter isolates in Italy. Sensors 14, 3308–3322. Foster, G., Holmes, B., Steigerwalt, A.G., Lawson, P.A., Thorne, P., Byrer, D.E., 2004. C. insulaenigrae sp. Nov., isolated from marine mammals. International Journal System Evolution Microbiology 54, 2369-2373. Friedman, C.R., Neimann, J., Wegener, H.C., Tauxe, R.V., 2000. Epidemiology of C. jejuni infections in the United States and other industrialized nations. In: Campylobacter, Second Ed. ASM Press, Washington DC, USA. pp. 121–138. Gannon, V.P.J., 1999. Control of Escherichia coli O157 at slaughter. In: Escherichia coli O157 in Farm Animals. Second Ed. Wallingford: CAB International, pp. 169–193. Girgis, S.S., Rashad, H.B., Othman, H.H., Bassim, N.N., Kassem, M.E., 2014. Multiplex PCR for Identification and Differentiation of Campylobacter Species and their Antimicrobial Susceptibility Pattern in Egyptian Patients. International Journal of current Microbiology and Applied Science 3, 861-875. Gonzalez, I., Grant, K.A., Richardson, P.T., Park, S.F., Collins, M.D., 1997. Specific identification of the entero-pathogens C. jejuni and C. coli using a PCR test based on the ceuE gene encoding a putative virulence determinant. Journal Clinical Microbiology 35, 759-63. Horrocks, S.M., Anderson, R.C., Nisbet, D.J., Ricks, S.C., 2009. Incidence and ecology of C. jejuni and coli in animals. Food Microbiology 15, 18-25. Kaakoush, N.O., Natalia, C., Hazel, M.M., Si Ming, M., 2015. Global Epidemiology of Campylobacter Infection. Clinical Microbiology Review 28(3), 687–720. Leuchtefeld, N.W., Wang, W.L.L., 1982. Animal reservoirs of C. jejuni. In: Campylobacter: Epidemiology. Pathogenesis and Biochemistry. Newell, D.G. (Ed.), MTP Press Ltd., Lancaster, England, pp. 249-52. Linton, D., Lawson, A.J., Owen, R.J., Stanley, J., 1997. PCR detection to species level and fingerprinting of C. jejuni and C. coli direct from diarrheic samples. Journal of Clinical Microbiology 35, 2568- 2572. Man, S.M., 2011. The clinical importance of emerging Campylobacter species. Nat Rev Gastroenterol Hepatol. 8, 669–685. Messens, W., Herman, L.D.E., Zutter, L., Heyndrickx, M., 2009. Multiple typing for the epidemiological study of contamination of broilers with Thermotolerant Campylobacter. Veterinary Microbiology 138, 120-131. Mdegela, R.H., Laurence, K., Jacob, P., Nonga, H.E., 2011. Occurrences of thermophilic Campylobacter in pigs slaughtered at Morocco slaughter slabs, Tanzania. Tropical Animal Health Production 43, 83–87. Moustafa, A.F.N., Ahmed, S.O., Ibrahim, A.A. Mosa, H.A., 2018. Prevalence of Zoonotic Species of Campylobacter in broiler Chicken and Humans in Assiut Governorate, Egypt. Approaches Poultry Dairy and Veterinary Science 3, 1-9. Moore, J.E., Corcoran, J.S., Dooley, S., Fanning, B., Lucey, M., Matsuda, D.A., Mcdowell, F., Mégraud, B., 2005. Campylobacter. Veterinary Research 36, 351-382. Ngulukun, S.S., 2017. Features, Detection, and Prevention of Foodborne Disease, In: Taxonomy and Physiological Characteristics of Campylobacter spp. Academic press, pp. 41-60. Nielsen, E.M., 2002. Occurrence and strain diversity of thermophilic Campylobacters in cattle of different age groups in dairy herds. Letters in Applied Microbiology 35, 85–89. On, S.L.W., 1996. Identification Methods for Campylobacter, Helicobacter, and related organisms. American Society for Microbiology Journal 9, 405–422. Oyofo, B.A., Thornton, S.A., Burr, D.H., Trust, T.J., Pavlovskis, O.R., Guerry, P., 1992. Specific detection of C. jejuni and C. coli by using polymerase chain reaction. Journal of Clinical Microbiology 30, 2613-2619. Peterson, M.C., 1994. Clinical aspects of Campylobacter jejuni infections in adults. West Journal of Medicine 161, 148–52. Presson, S., Olsen, K.E., 2005. Multiplex PCR for identification of C. coli and C. jejuni from pure cultures and directly on stool samples. Journal of Medical Microbiology 54, 1043-1047. Rautelin, H., Jusufovic, J., Hänninen, M.L., 1999. Identification of hippurate-negative thermophilic Campylobacters. Diagnostic Microbiol. Infect. Dis., 35, 9-12. Roberts, D., Greenwood, M., 2003. Isolation and enrichment of microorganisms, In: Practical Food Microbiology, third Ed. D. Blackwell Publishing Ltd., Malden, MA, pp. 131–192. Ruiz-Palacios, G.M., 2007. The health burden of Campylobacter infection and the impact of antimicrobial resistance: playing chicken, “Clinical Infectious Diseases 44, 701–703. Sails, A.D., Bolton, F.J., Fox, A.J., Wareng, D.R.A., Greenway, D.L.A., 1998. A reverse transcriptase polymerase chain reaction assay for the detection of thermophilic Campylobacter Spp. Mol. Cell. Prob.12, 317-322. Shimaa, T., Omara, H.A., El Fadaly, A., Barakat, M.A., 2015. Public Health Hazard of Zoonotic C. jejuni Reference to Egyptian Regional and Seasonal Variations. Research Journal of Microbiology 10, 343-354. Silva, J.D., Leite, M., Fernandes, C., M, ena P.A., Gibbs, P., 2011. Campylobacter spp. as a foodborne pathogen: a review. Frontier Microbiology 2, 200-212. Skirrow, M.B., Blaser, M.J., 2000. Clinical aspects of Campylobacter infection. In: Campylobacter, Second Ed. ASM Press, Washington DC, USA, pp 69–88. Stanley, K.N., Wallace, J.S., Currie, J., Diggle, P., Jones, K., 1998. The seasonal variation of thermophilic Campylobacters in lambs at slaughter. Journal of Applied Microbiology 84, 1111–1116. Stanley, K., Jones, K., 2003. Cattle and sheep farms as reservoirs of Campylobacter. Journal of Applied Microbiology 94, 104−111. Stucki, U., Frey, J., Nicolet, J., Burnens, A.P., 1995. Identification of C. jejuni on the basis of a species-specific gene that encodes a membrane protein. Journal Clinical Microbiology 33, 855-9. Workman, S.N., Sobers, S.J., Mathison, G.E., Lavoie, M.C., 2006. Human Campylobacter-associated enteritis on the caribbean island of barbados. American Journal Tropical Medical Hygiene 74, 623-627. Van Eys, G.M., Gravekamp, C., Gerritsen, M., Quint, W., Cornelissen, M., Schegges, J.W., 1989. Detection of leptospires in urine by polymerase chain raction. Journal Clinical Microbiology 27, 2258–2262. Van Deun, K., Haesebrouck, F., Hendrickx, M., Favoreel, H., Dewulf, J., Ceelen, L., Dumez, L., Messens, W., Leleu, S., Van Immersal, F., Ducatelle, R., Pasmans, F. 2007. Virulence properties of C. jejuni isolates of poultry and human origin. Journal Medical Microbiology 56, 1284–1289. WHO, 2017. World Health Organization: Campylobacter. http:// WHO.Org. |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||